🚀 Getting Started with IDUN on Windows
IDUN is NTNU’s high-performance computing (HPC) cluster, designed for large-scale simulations, optimization, and data-intensive research.
For Windows users, the workflow typically includes:
- 📁 Accessing IDUN storage (network drive)
- 🔐 Connecting via SSH (PuTTY)
- ⚡ Running jobs (interactive / SLURM)
- 🧪 Managing environments (Conda, Python)
- 📊 Monitoring resources
This guide walks you through a practical, step-by-step setup.
📁 1. Access IDUN Files from Windows (Recommended)
The easiest way to work with IDUN is by mounting it as a network drive.
⚠️ Requires NTNU VPN or campus network
Steps
1. Open File Explorer → This PC
2. Click Map Network Drive
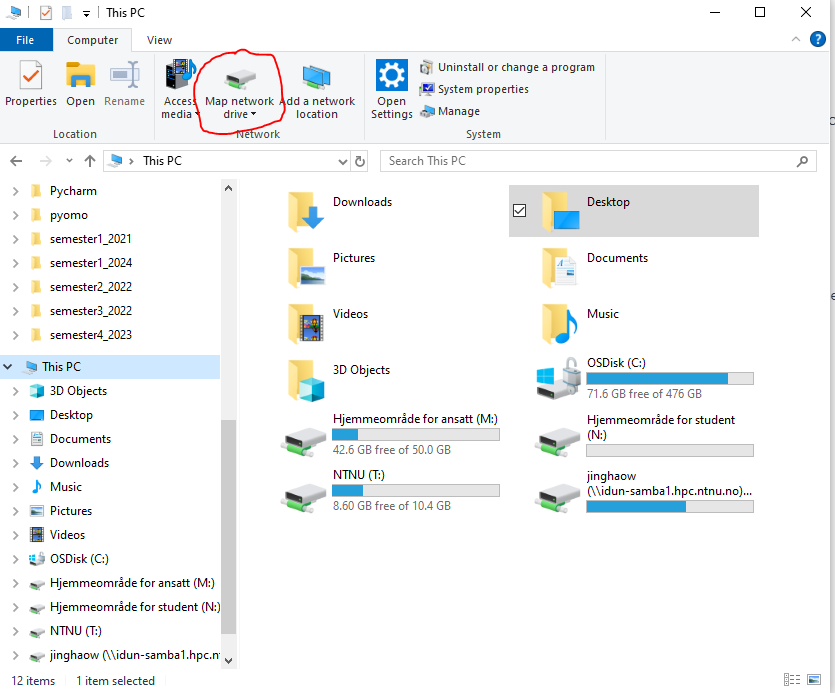
3. Enter network path
1
| \\idun-samba1.hpc.ntnu.no\<username>
|
<username> = your NTNU FEIDE username - Tick: ✅ Connect using different credentials
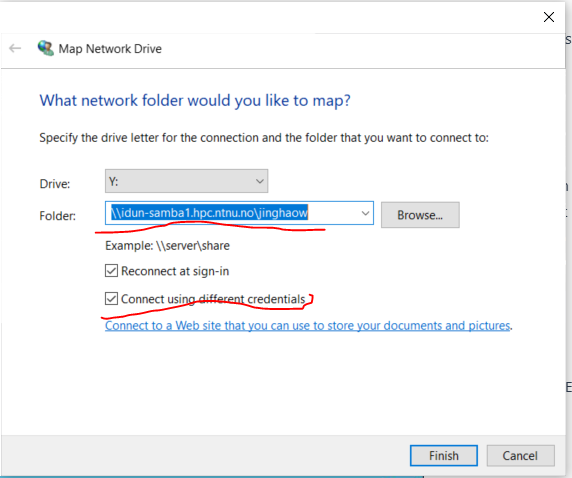
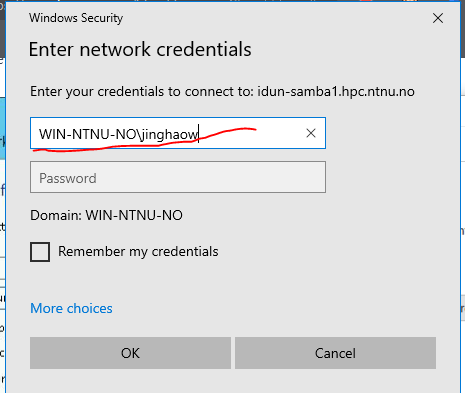
5. Enter password → Done ✅
You now have direct access to IDUN storage from Windows.
📚 More info:
https://www.hpc.ntnu.no/idun/documentation/transferring-data/
🔐 2. SSH Login via PuTTY (Optional)
PuTTY allows terminal access to IDUN.
or
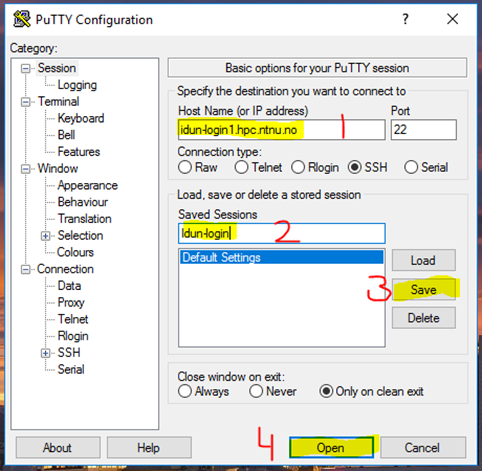
2. Login
- Username: NTNU username
- Password: (hidden input)
⚡ 3. Request Compute Resources
You must request compute resources before running jobs.
Interactive session
1
| salloc --account=share-ie-iel --nodes=1 --cpus-per-task=16 --time=01:00:00 --constraint="pec6520&56c"
|
Explanation
salloc → allocate resources --account → project account --nodes → number of nodes --cpus-per-task → CPU cores --time → max runtime --constraint → hardware selection
Check available resources
1
| sinfo -a -o "%22N %8c %10m %30f %10G"
|
Connect to assigned node
🌐 4. Using OnDemand (Web Interface)
IDUN provides a modern web interface:
👉 https://apps.hpc.ntnu.no
Login → Clusters
🧪 5. Python & Conda Environment
Load Anaconda
1
| module load Anaconda3/2022.10
|
Create environment
1
2
| conda create -n <env_name> python=3.8 --channel conda-forge --yes
conda activate <env_name>
|
⚠️ Important: Always load Anaconda before using Conda
1
| module load Anaconda3/2022.10
|
Add Gurobi channel
1
2
| conda config --add channels https://conda.anaconda.org/gurobi
conda config --show channels
|
Install Gurobi
1
| conda install gurobi=9 --yes
|
Add conda-forge (recommended)
1
2
| conda config --env --add channels conda-forge
conda config --env --set channel_priority strict
|
Install Pyomo & solvers
1
2
3
| conda install openpyxl
conda install -c conda-forge pyomo
conda install -c conda-forge ipopt glpk
|
📓 7. Jupyter Notebook Setup
Install kernel
1
2
| conda install ipykernel
ipython kernel install --user --name=<env_name>
|
Check kernels
⚙️ 8. SLURM Jobs (Batch Processing)
Sequential job (sbatch)
Example Python file
1
2
| for i in range(20):
print(i)
|
SLURM script
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
| #!/bin/bash
source /cluster/apps/eb/software/Anaconda3/2022.10/etc/profile.d/conda.sh
conda activate <env_name>
python test.py
|
Submit job
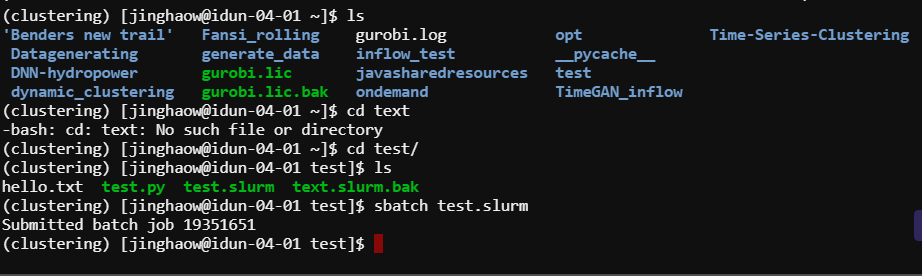
Output example
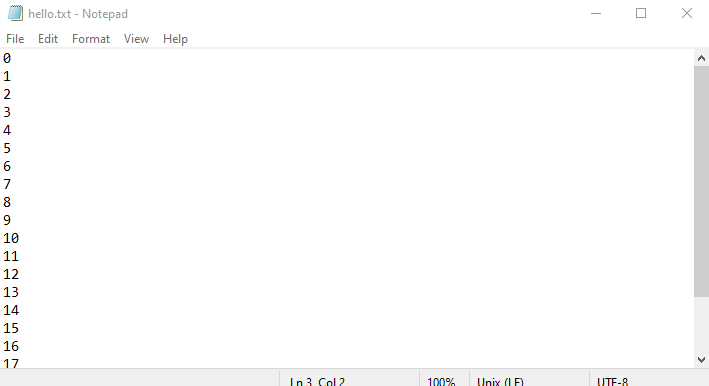
⚡ 9. Parallel Jobs (srun)
Python example
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
| def test(numb):
return numb
def run_test(index):
state_index = (index-1)*10
end_index = index*10
DATA = {}
for a in range(state_index, end_index):
DATA[a] = test(a)
with open(f'output/data_output{index}.txt', 'w') as f:
f.write(str(DATA))
if __name__ == '__main__':
import sys
index = int(sys.argv[1])
run_test(index)
|
SLURM script
Logs & output
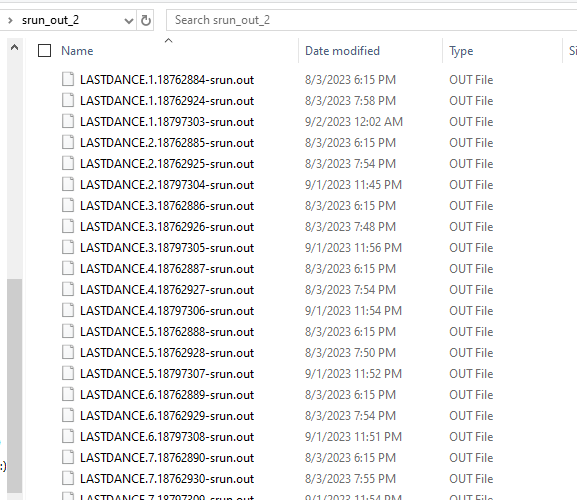
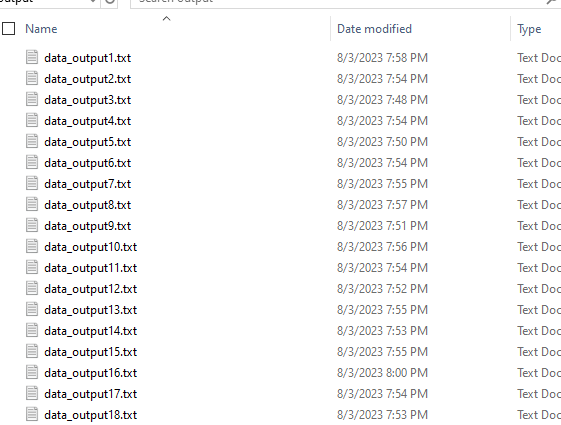
🛠️ Useful Commands
Conda
1
2
| conda list
conda remove <package>
|
System
Edit .bashrc
i → insert Esc + :wq → save Esc + :q → quit
Check resources
1
| sinfo -a -o "%22N %8c %10m %30f %10G"
|
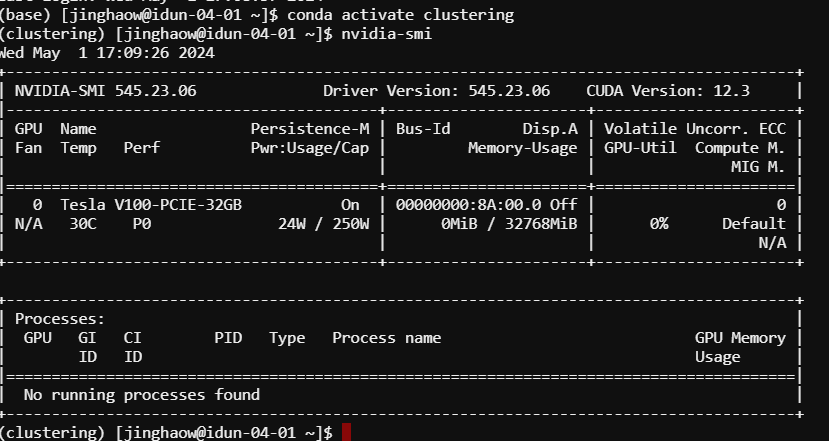
Monitor usage
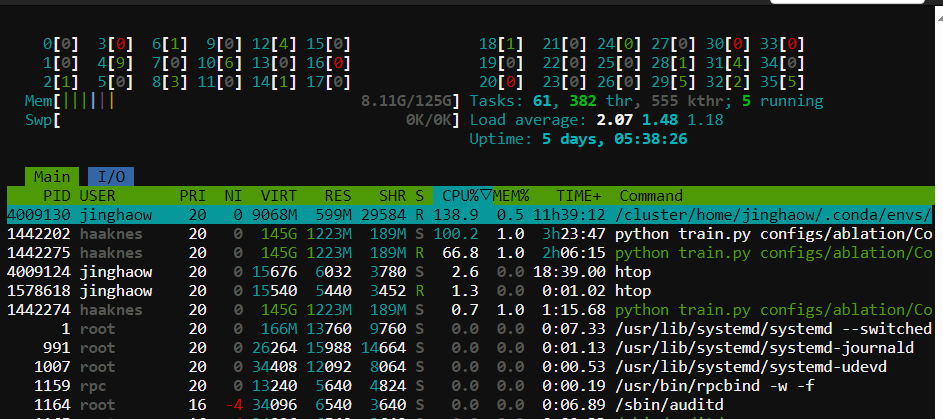
📚 Additional Resources
🧠 Final Notes
- Use network drive for easy file transfer
- Use SLURM (sbatch/srun) for real workloads
- Use Conda environments for reproducibility
- Prefer OnDemand UI if you’re new
🙏 Acknowledgement
Special thanks to Runar Mellerud and Anders Gytri for support and documentation on IDUN usage.
–Powered by automatic agent: OpenAI